As a Bioinformatician, often I get to work with PDX cancer samples. I’ve recently been reading about samples containing genome admixture, and was revisiting strategies that we commonly use for analyzing these biological data. Presented here is a summary of the existing software tools used for this purpose.
Continue readingThe ANCHOR pipeline
ANCHOR is a high-resolution metagenomics pipeline, the result of a multi-disciplinary collaboration between C3G and researchers at Institut de recherche en biologie végétale (IRBV) at Université de Montreal. Published in 2019 in the journal Environmental Microbiology, the pipeline demonstrated unprecedented accuracy in its characterization of microbiome samples.
Continue readingCharacterizing the landscape of genetic drivers and their clinical impact
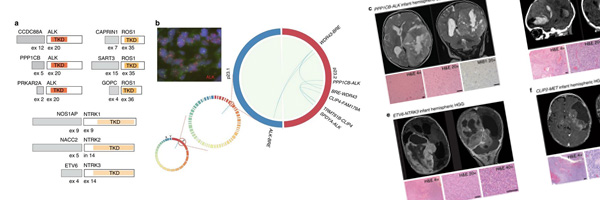
Recently published in [Nature Communications], this paper presents work by members of the C3G Toronto node that integrates genomic and transcriptomic analyses to assess the molecular and clinical features of infant glioma patients.
Continue readingFibromyalgia
This summer has also been very special for C3G’s metagenome specialist Emmanuel Gonzalez with a publication in Pain highlighting a strong potential link between the microbiome and fibromyalgia, a terrible and elusive disorder affecting a large fraction of the population.
Continue readingGSoC 2019
Again this year, C3G was a Google Summer of Code organization. For people unfamiliar with it, GSoC is in Google’s own words: ” a global program that matches students up with open source, free software and technology-related organizations to write code and get paid to do it! ”
Continue readingAltered differentiation is central to HIV-specific CD4
Another noteworthy publication to which C3G members contributed as authors was published in Nature Immunology this summer.
Continue readingMethylation signatures investigations
The C3G Toronto node has also been involved in two projects to investigate methylation signatures for specific conditions.
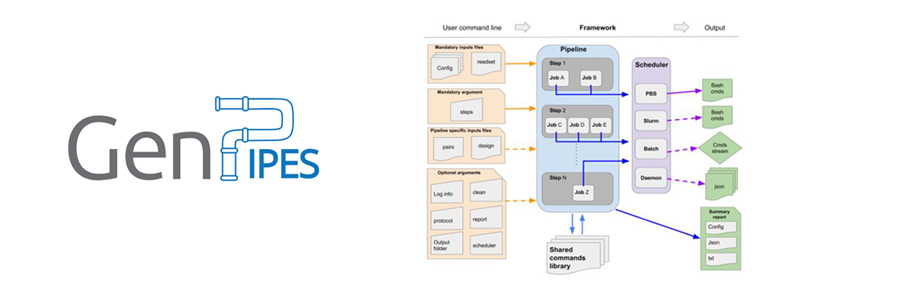
Continue readingGenPipes: an open-source framework
It started in June with the publication of our beloved GenPipes framework and set of NGS data analysis pipelines in GigaScience.
Continue readingCovid-19 updates at C3G
While our host institutions (McGill University, SickKids Toronto) have put various measures in place to ensure the safety of our staff, our platform remains fully operational and available to support genomics research.
Continue readingForCasT
ForCas Tool (ForCasT) is a comprehensive tool for the design, evaluation and collection of CRISPR/Cas9 guide RNAs and primers. Using robust parameters, it generates guide RNAs for target loci, assesses their quality for any potential off-target effects and designs associated primers.
Continue reading