Omics Analysis Services
C3Gs team of experts in bioinformatics and software development offer extensive experience to analyze complex ‘omics data and transform it significant discoveries. Our teams offers wide range of experience in a wide range of ’omic analysis, leveraging state of the art computational methods.
Our services encompass:
Personalized data analysis by expert bioinformaticians
Custom bioinformatics software development
Databases and portals for data organization and sharing
Experimental design support
Collaboration on grant applications
On-demand bioinformatics training
Portfolio
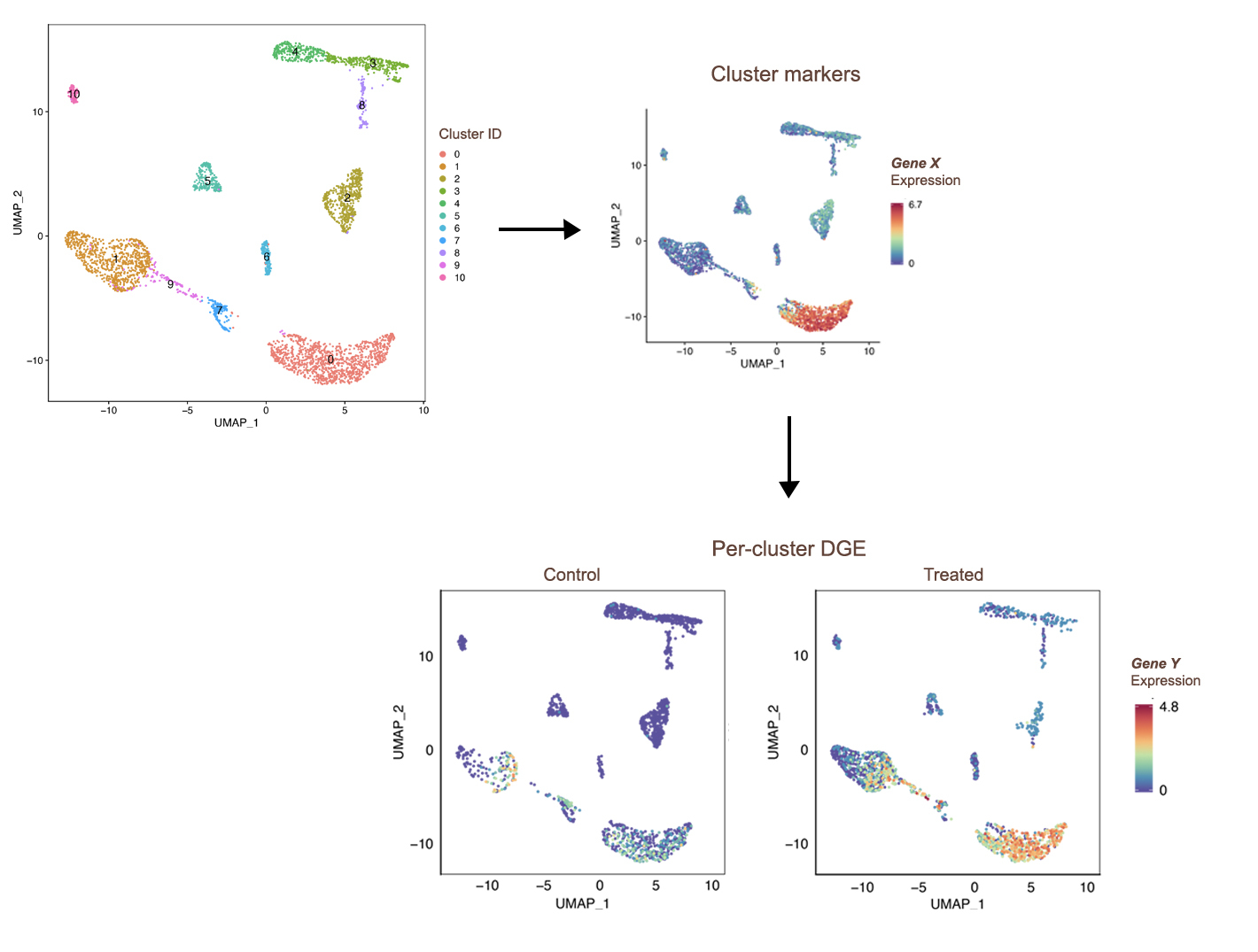
Single-cell Expression
Single-cell RNA sequencing (scRNA-seq) has greatly expanded the study complex biological systems by enabling genome-wide expression profiling at the level of individual cells. We offer comprehensive scRNA-seq analysis services, including:
- Raw Data Processing: Alignment and generation of gene-barcode matrices using Cell Ranger, alevin fry, or equivalent tools.
- Quality Control & Filtering: Rigorous assessment of cell quality metrics, including detection of low-quality cells, doublets, and ambient RNA contamination
- Demultiplexing: Genetic and algorithm based sample demultiplexing from pooled single cell libraries.
- Normalization & Correction: Data normalization, batch correction, and ambient RNA correction and other steps to ensure accurate downstream analysis.
- Clustering & Cell Type Identification: Clustering of cells and annotation of cell types based on marker gene expression.
- Data Exploration: Visualization and exploration tools to support hypothesis generation.
- Differential Expression & Abundance Analysis: Identification of genes and cell populations with differential expression or abundance between conditions or groups.
- Trajectory & Pseudotime Analysis: Reconstruction of dynamic cellular processes such as differentiation pathways and lineage relationships.
- Multi-Omic Integration: Integration with complementary data types, including single-nucleus ATAC-seq (snATAC-seq), immune profiling (e.g., CITE-seq), and spatial transcriptomics.
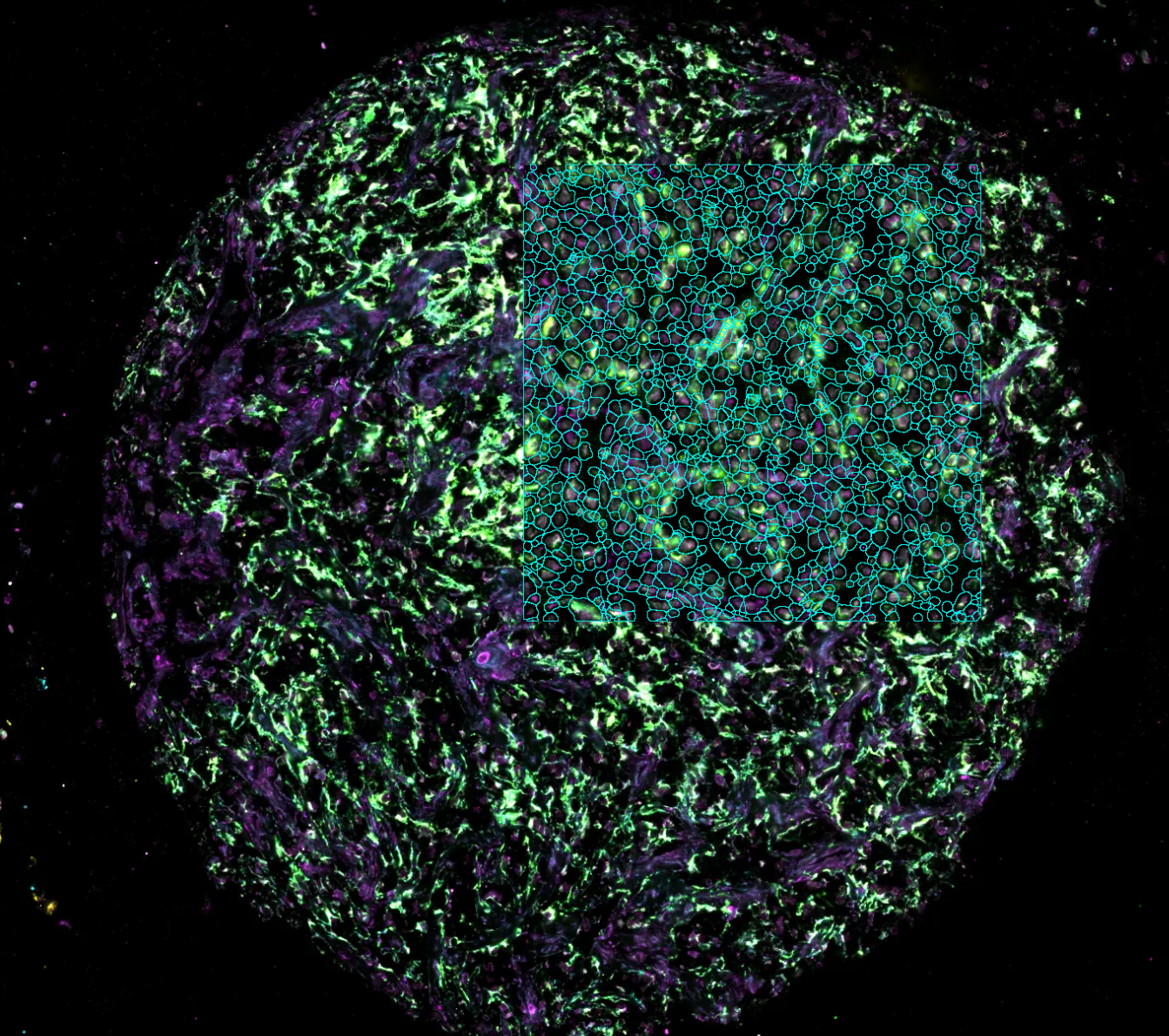
Spatial Transcriptomics
Mapping gene expression within its original spatial context in a tissue. Spatial transcriptomics publications have increased exponentially since 2018, it is highly used in fields like oncology, neuroscience, immunology, and developmental biology. We offer comprehensive analysis services for varied technologies like Visium or GeoMx, including:
- Raw Data Processing: Alignment and generation of gene-barcode matrices
- Image processing: Cell segmentation and transcript – coordinate identification.
- Quality Control & Filtering: Rigorous assessment of cell quality metrics, including detection of low-quality cells
- Normalization & Correction: Data normalization, batch correction, and ambient RNA correction and other steps to ensure accurate downstream analysis.
- Clustering & Cell Type Identification: Clustering of cells and annotation of cell types based on marker gene expression.
- Data Exploration: Visualization and exploration tools to support hypothesis generation.
- Differential Expression & Abundance Analysis: Identification of genes and cell populations with differential expression or abundance between conditions or groups.
- Gene expression and tissue mapping: Reconstruction of dynamic cellular processes such as differentiation pathways and lineage relationships.
- Multi-Omic Integration: Integration with complementary data types, including single-nucleus ATAC-seq (snATAC-seq), immune profiling (e.g., CITE-seq), and spatial transcriptomics.
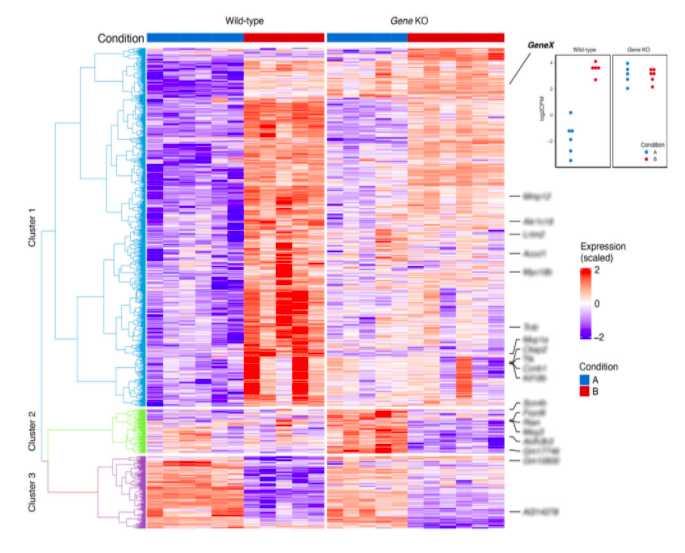
RNA-Seq
As one of the most commonly requested services, our Services team can help you get the most out of RNA-seq data. This service typically includes:
- Preprocessing: Alignment, assembly, quantification of expression
- Quality Control & Filtering: Rigorous assessment of sample quality metrics, including detection of low count genes, exploratory data analysis (EDA), Principal component analysis (PCA), etc.
- Differential expression & pathway analysis: Downstream analysis of transcript abundance, including differential expression, Gene Ontology (GO), and KEGG pathway enrichment analysis.
- Customized visualisations.
- Gene/Pathway analysis: gene set expression (GSEA) & Gene Ontology (GO) analyses & Pathway enrichment
- Multi-Omic Integration: Integration of RNA-seq data with complementary data types, including scRNA-seq, ChIP-seq, ATAC-seq, etc.
- Other RNA based analysis: Alternate-splicing analysis, eSNV calling, fusion detection, deconvolution, PDX, decontamination etc.
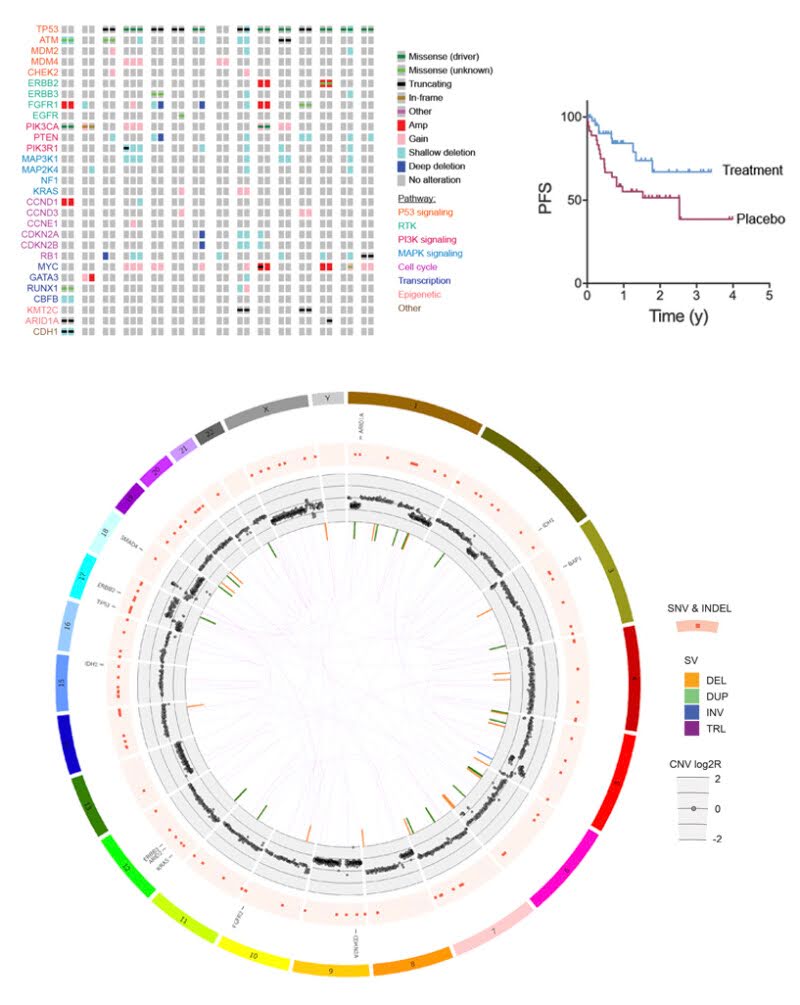
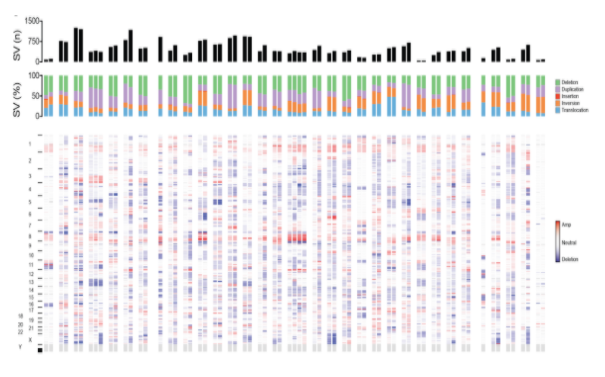
Cancer Genomics
By sequencing DNA from both tumour and matched normal (healthy) samples, whole-genome sequencing (WGS) enables the comprehensive discovery of novel cancer-associated variants. Our cancer bioinformatics services provide an end-to-end analytical solution, including:
- Somatic variant detection: Identification of somatic SNVs, indels, copy number alterations, and structural variants using an ensemble approach combining different tools.
- Clinical and functional annotation: Integration of cancer-relevant annotations using Personal Cancer Genome Reporter (PCGR) and Cancer Predisposition Sequencing Reporter (CPSR), providing insights into functional impact, treatment relevance, mutational burden, and microsatellite instability (MSI).
- Structural variant analysis: Application of GRIDSS–PURPLE–LINX for high-resolution detection and interpretation of somatic structural variation, combining SV calls, copy-number profiling, and event-level annotation to elucidate complex genomic rearrangements.
- Quality control and sample identity verification: Rigorous QC pipeline and cross-sample identity checks to ensure data integrity and sample concordance.
- Multi-Omic Integration: Integration of matched RNA-seq data for the identification of expressed variants and gene fusions, enhancing biological and clinical interpretation.
- Support for model systems: Tailored workflows for patient-derived xenograft (PDX) and mouse tumour models.
- Rapid analysis for prospective studies: Fast-track options available to support prospective clinical and translational studies.
Whole Genome Sequencing & Exome-Seq
We have extensive experience in identifying variants from whole-genome sequencing (WGS) or whole-exome sequencing (WES) data. This service generally involves:
- Processing raw sequencing data to variant calls following the GATK best practices
- Detecting structural variants and CNVs
- Annotating and filtering variants according to specific needs of the project to help with variant prioritization (e.g. de novo, compound het. in trios using the GEMINI/Slivar framework)
- Genome-Wide Association Study (GWAS)
- Multi-Omic Integration
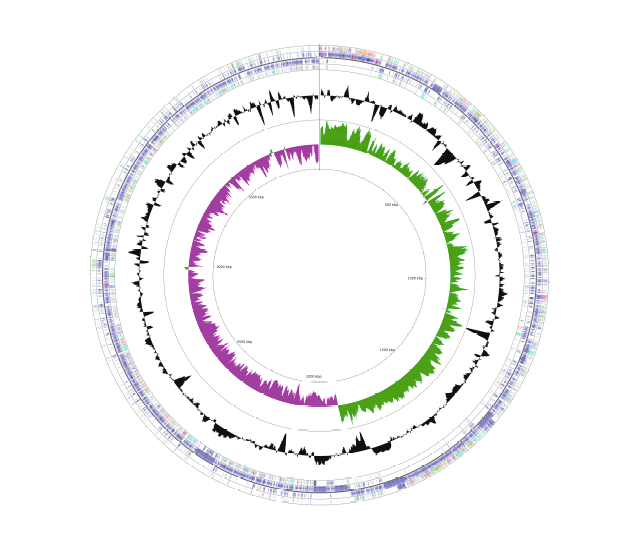
Genome Assembly & Annotation
Long-reads sequencing (eg. PacBio Hi-Fi, ONT) allows for large and difficult regions of the genome (eg. TE’s, repeats, rRNA islands) to be captured in single long reads which can then be accurately anchored in the de novo assembly. We have extensive experience in de novo assembling small (haploid; prokaryotes, archaea) and large (diploid; eukaryotes) novel genomes using both long read technologies and short reads for hybrid assembly approaches. Our toolbox includes:
- Assembly and refinement of de novo assembly using data from supporting technologies
- Complete annotation processes: For example using PGAP. BRAKER4, EGAPX
- Detection, polishing and circularization of bacterial chromosome/s or other circular molecules (plasmid, mitochondria)
- Comparative genomics with existing assemblies: For example using OrthoFinder
- Reference-based variant detection: INDEL, SNP, large structural rearrangements, phased alleles
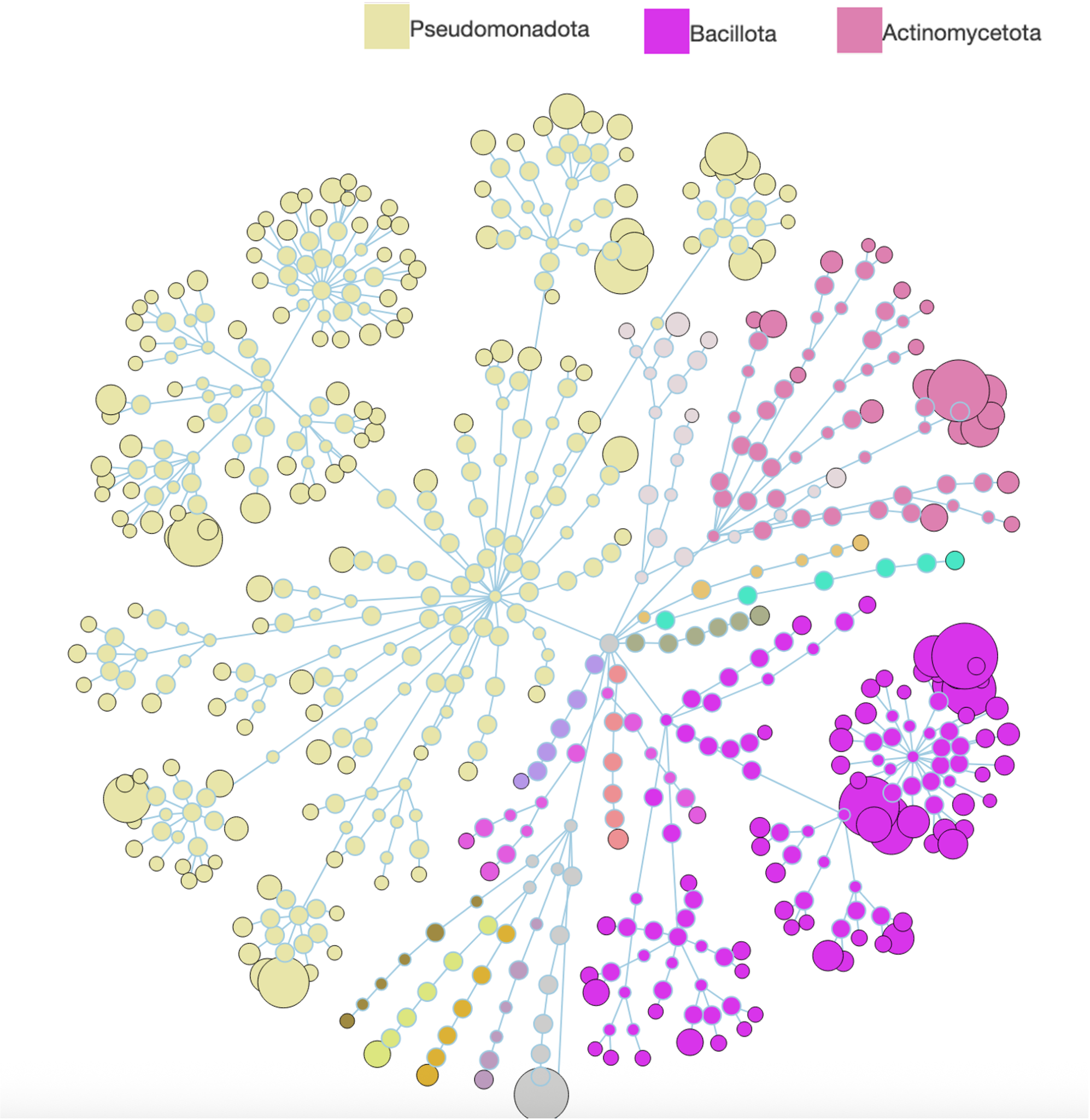
Microbiome & Metagenomics
We are committed to providing top-tier services for metagenomics and metatranscriptomics research. Our state-of-the-art, fully integrated analytical pipelines and expertise support a wide range of study designs, including:
- Amplicon-based metagenomics (16S, 18S, ITS, COI)
- Shotgun metagenomics (WGS)
- Metatranscriptomics (RNA-Seq)
Our analyses enable advanced insights such as KEGG global metabolic networks and co-occurrence networks, which can be generated from WGS and 16S outputs using the MicrobiomeAnalyst interactive platform.
Core deliverables for amplicon-based metagenomics (16S, 18S, ITS, COI) include:
- Denoising of raw reads to ASVs using a DADA2- or QIIME-based workflow
- Taxonomic assignment of ASVs
- Functional annotation of the microbial community
- Output files fully compatible with MicrobiomeAnalyst and Calypso, enabling researchers to create customized, publication-ready figures with ease.
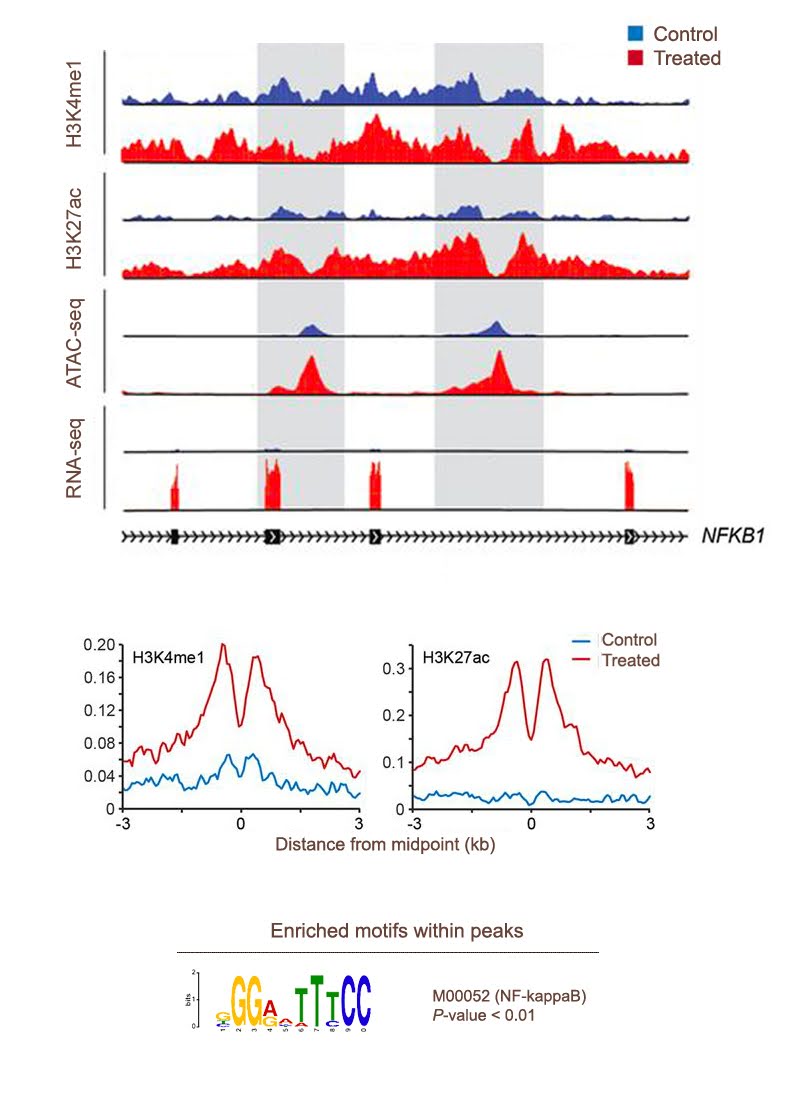
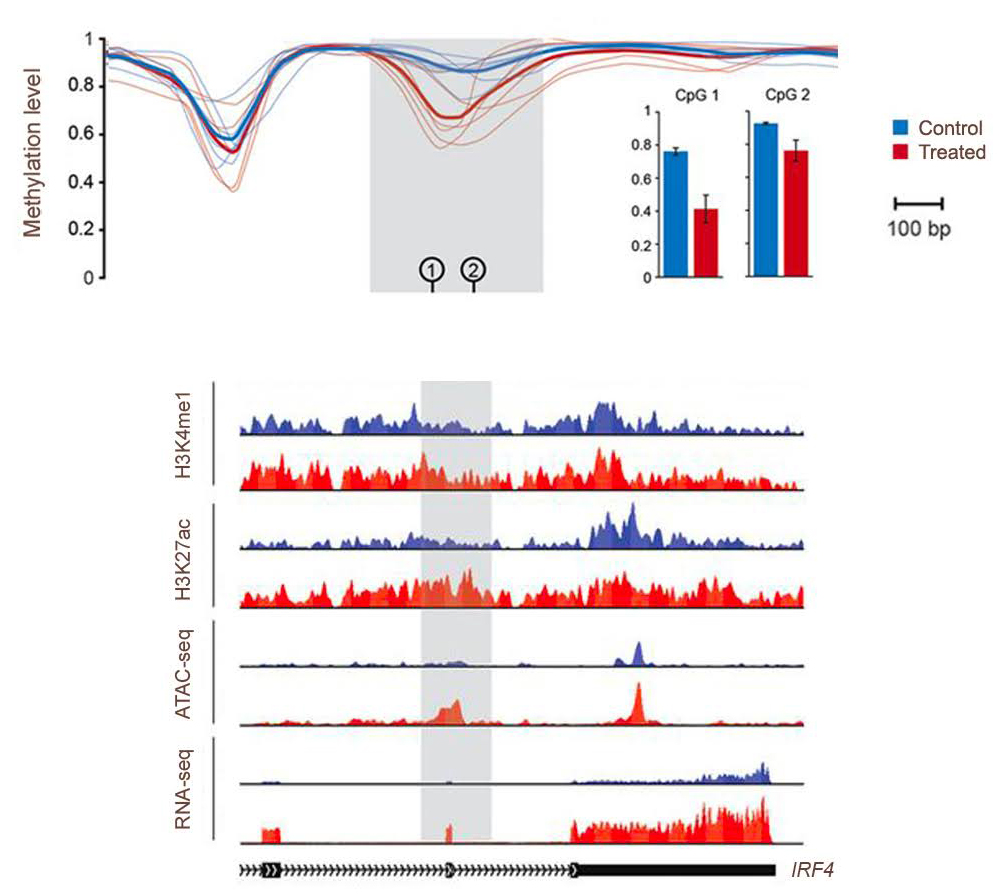
Epigenomic Bioinformatics
Genome-wide profiles of DNA-protein interaction/histone modification and chromatin accessibility sites provide valuable insights into the regulatory genome landscape.
- Peak Calling and Annotation: For ChIP-seq, ATAC-seq and similar assays (e.g. CUT&RUN), we deliver a list of significant peaks that are annotated with genomic context as well as their motif enrichments.
- Genome Coverage: For Methyl-seq, alignment of bisulphite-converted reads to a reference genome and the quantification of methylation levels.
- Genome-Wide chromatin interaction: For Hi-C, construction of contact matrices and generation of multi-resolution contact maps for downstream interpretation followed by Topologically Associating Domains (TADs) identification and annotation.
- Custom visualization: We generate the epigenetic tracks via UCSC Genome Browser or WashU epigenome browser. Metagene plots and enrichment density heatmaps are also typically performed.
- Differential Analysis: reveal peaks (Or genomic regions) with statistically significant signal variation across treatments or conditions.
- Multi-Omic Integration: Integration with complementary data types, including scRNA-seq, RNA-seq, ChIPseq, ATAC-seq, Methyl-seq, HiC, etc
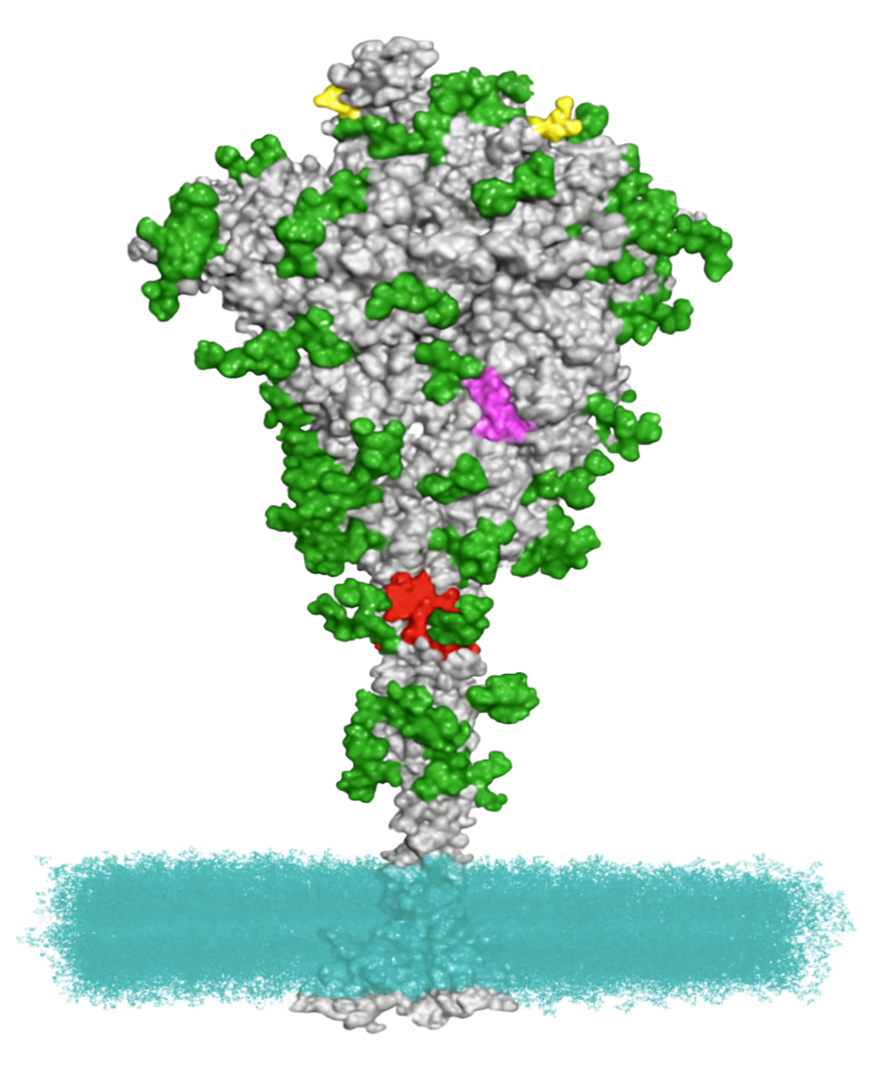
Structural Bioinformatics
We can help you with structural bioinformatics that integrates with other genomics studies. This service typically includes:
- Structure visualization
- Mutation mapping
- MD simulation, binding energy
- Contact maps
De novo Transcriptome (RNA-seq) Assembly
For organisms lacking a high-quality reference genome or transcriptome, our de novo RNA-seq assembly services enable comprehensive characterization of the transcriptome using cutting-edge approaches. Our services include:
- Transcriptome assembly & quantification: De novo assembly of RNA-seq data using Trinity, followed by transcript abundance estimation and quality assessment.
- Functional annotation: Comprehensive transcript annotation using Trinotate, incorporating homology searches, protein domain identification, and functional classification.
- Differential expression & pathway analysis: Downstream analysis of transcript abundance, including differential expression, Gene Ontology (GO), and KEGG pathway enrichment analysis.
- Visualization & reporting: Generation of customized visualizations and reports to facilitate the interpretation and communication of results.
- Multi-Omic Integration: Integration of RNA-seq data with complementary data types, including scRNA-seq, ChIP-seq, ATAC-seq, etc
We provide flexible, project-specific support for a wide range of research applications, from basic biology to applied genomics in non-model organisms.
Other – Omics
- Other Single-Cell Assays (snATAC-seq, snDNA-seq)
- Expression & Methylation microarrays
- Misc. sequencing assays: DRIPc-seq, RIP-seq, etc.
- miRNA and other small RNA-seqs
- CRISPR-Cas9 data and statistical analysis
- Sequencing of pools (Pool-seq)
- Population Genomics: GBS, RAD-Seq
- CyTOF
- Processed Metabolomics data integration.
- Processed Proteomics data analysis.
- Integrative Analyses – ex Similarity Network Fusion (SNF)
- Network Visualizations and Analysis
- Image Analysis
Testimonials