Recently published in [Nature Communications], this paper presents work by members of the C3G Toronto node that integrates genomic and transcriptomic analyses to assess the molecular and clinical features of infant glioma patients. Examining single nucleotide variants, changes in copy number, fusion formation and other transcriptomic analyses revealed three clinical glioma subgroups in infants, each with distinct genetic drivers, locations in the brain and responses to treatment. Gliomas in infants have substantially different treatment outcomes compared to those that occur in children and adults, yet little is understood about the molecular basis of these differences. This paper gives a comprehensive molecular analysis of infant gliomas to helps to ascertain the biological mechanisms driving their oncogenesis and to help guide future diagnostics and treatment approaches for these patients.
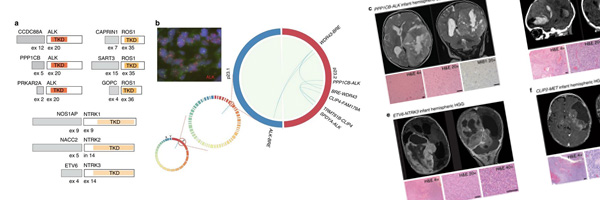
Characterizing the landscape of genetic drivers and their clinical impact
Share this post on social media:
Facebook
X
Linkedin
Recent Posts
- Inside GenPipes: Exploring C3G’s Software Solution for Life Science Research
- C3G Receives $4.6 million in Funding from the Canadian Foundation for Innovation (CFI) and partner organizations
- A beginner’s Guide to Manual Curation of Transposable Elements
- C3G Bioinformatics Platform at the Rosalind and Morris Goodman Cancer Institute
- The SecureData4Health (SD4H) website is LIVE
- Gene fusion meta-calling with MetaFusion
- Working with Genomics Coming From a Pure Software Engineering Background
- Understanding why persons living with HIV are at increased risk of tuberculosis
