Designing a sequencing project? A recent Genome Biology study offers a practical framework for choosing the right technologies based on your budget and research objectives.
Accurately identifying every genetic variation in the human genome is essential for both research and clinical applications. This Genome Biology study brings together expertise from the Canadian Centre for Computational Genomics (C3G) (Robert Eveleigh, Jose Hector Galvez, Mathieu Bourgey, and Guillaume Bourque) and the Advanced Genomic Technologies Laboratory (Sarah Reiling and Jiannis Ragoussis) to benchmark state-of-the-art sequencing platforms and variant detection approaches across both small and large variant classes.
Methodology
Using the Genome in a Bottle (GIAB) HG002 reference sample, the team systematically compared short-read (SRS; Illumina, MGI) and long-read (LRS; PacBio Sequel/Revio, ONT R9/R10) technologies against Telomere-to-Telomere (T2T) and Clinically Medically Relevant Genes (CMRG) benchmarks, spanning a range of sequencing depths, genomic contexts, and bioinformatic pipelines.
Findings
Their findings confirm that platform and workflow selection should be driven by research objectives. SRS excels at identifying small variants in well-mapped regions, while LR, particularly PacBio Revio, delivers superior accuracy for structural variants and small variants in complex or repetitive regions, achieving accuracy saturation at markedly lower sequencing depths (20–45×) than short-reads (>60×).
While SRS remains a practical choice for high-throughput genotyping, LRS provides the resolution required for clinical applications at challenging loci.
Learn More
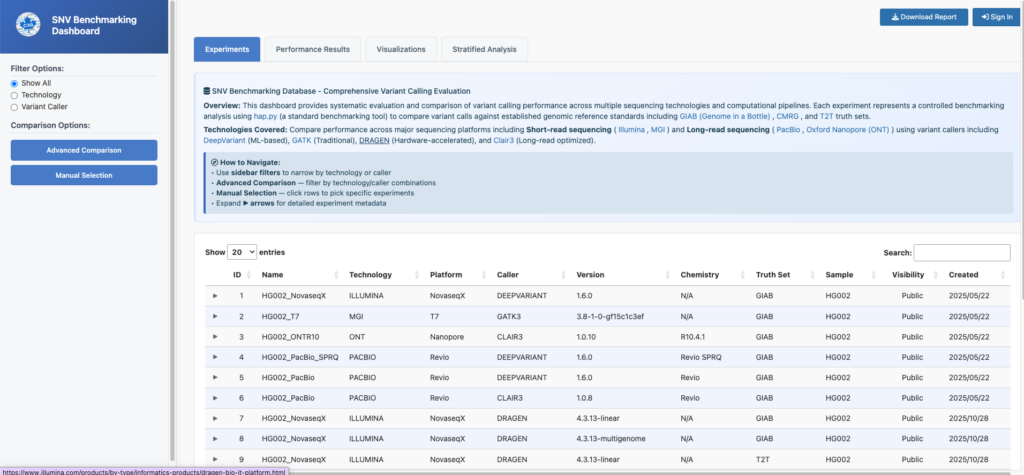
To stay current with the latest algorithms, technologies, and benchmarking practices, C3G also maintains a live dashboard!